COMPARATIVE CONSERVATION OF MEIOTIC PROTEINS IN DIFFERENT PHYLOGENETIC LINES OF EUKARYOTES
DOI: https://doi.org/10.18454/jbg.2018.2.7.1
Grishaeva T.M.*
Department of Cytogenetics, Vavilov Institute of General Genetics, Russian Academy of Sciences, Moscow, Russian Federation
* Correspodning author (grishaeva@vigg.ru)
Received: 18.03.2018; Accepted: 05.04.2018; Published: 22.05.2018
COMPARATIVE CONSERVATION OF MEIOTIC PROTEINS IN DIFFERENT PHYLOGENETIC LINES OF EUKARYOTES
Research article
Abstract
Motivation: Meiosis — a two-stage process of sex cell division — is served by several hundreds of proteins. A part of them went to eukaryotes from prokaryotes, others appeared in first eukaryotes, and some proteins appeared de novo in multicellular eukaryotes. We compared the conservation of proteins involved in various processes occurring in meiosis.
Results: The conservations of five meiotic enzymes (MLH1, MRE11, MSH4, BRCA1, BRCA2) and three silencing markers (histone H2AX, SUMO1, ATR) were compared using a set of bioinformatics methods. Orthologs of these proteins from the proteomes of model species were compared, representing different lines of development of eukaryotes. Among the enzymes, the most conserved is MLH1, which provide correction of mismatch bases, and the least conserved are BRCA1 and BRCA2 repair enzymes which are present only in vertebrates. Among silencing proteins, histone H2AX is the most conserved one, playing the central part in the regulation of the transcription, in the repair and replication of DNA. The small protein SUMO1, which is involved in many cellular processes, is less conserved. ATR kinase in different species is similar only in the C-terminal part of the molecule.
Keywords: silencing of chromatin, meiosis, proteins, conservation.
Гришаева Т.М.*
Лаборатория цитогенетики, ФГБУН Институт общей генетики им. Н.И. Вавилова Российской Академии Наук, Москва, Россия
* Корреспондирующий автор (grishaeva@vigg.ru)
Получена: 18.03.2018; Принята: 05.04.2018; Опубликована: 22.05.2018
СРАВНИТЕЛЬНАЯ КОНСЕРВАТИВНОСТЬ МЕЙОТИЧЕСКИХ БЕЛКОВ В РАЗНЫХ ФИЛОГЕНЕТИЧЕСКИХ ЛИНИЯХ ЭУКАРИОТ
Научная статья
Аннотация
Мотивация: Мейоз – двухступенчатый процесс деления половых клеток –обслуживается несколькими сотнями белков. Часть из них досталась эукариотам от прокариот, другие появились у первых эукариот, а некоторые белки возникли de novo у многоклеточных эукариот. Мы сравнили консервативность белков, принимающих участие в разных процессах, происходящих в мейозе.
Результаты: Проведено сравнение консервативности пяти ферментов мейоза (MLH1, MRE11, MSH4, BRCA1, BRCA2) и трёх маркеров сайленсинга (гистон H2AX, SUMO1, ATR) с помощью комплекса методов биоинформатики. Сравнивали ортологи этих белков из протеомов модельных видов, представляющих разные линии развития эукариот. Среди ферментов наибольшей консервативностью обладает обеспечивающий коррекцию неправильно спаренных оснований, а наименее консервативны ферменты репарации BRCA1 и BRCA2, присутствующие только у позвоночных. Из белков сайленсинга наибольшей консервативностью отличается гистон H2AX, играющий центральную роль в регуляции транскрипции, в репарации и репликации ДНК. Менее консервативен небольшой белок SUMO1, принимающий участие во многих клеточных процессах. Киназа ATR у разных видов сходна только в С-концевой части молекулы.
Ключевые слова: сайленсинг хроматина, мейоз, белки, консервативность.
1. Introduction
Meiosis — a two-stage process of sex cell division, as a result of which haploid cells (gametes) are formed from a diploid one. During the first division of meiosis, the assortment of homologous chromosomes takes place, during the second one — the segregation of sister chromatids. In addition to the segregation, meiosis is accompanied by the recombination of genetic material. The meiosis process is served by more than a thousand proteins, some of which are specific for meiosis [10], [17]. The main mass of meiosis-specific proteins consists of recombination proteins — enzymes and modulators of this process. They are considered the most conserved [7], [9]. The mismatch-repair enzyme MLH1 is one of the universal participants in early events in the chain of molecular processes leading to the crossing over. This protein has the same function in all organisms, and it consists in the fact that MLH1 must control compliance with the rules of complementarity of nucleotides in the structure of DNA double helix [9]. The MSH4 protein belongs to the mutS family and is necessary for the reciprocal recombination (in the final stages) and for the correct segregation of homologous chromosomes in the first division of meiosis. It works in the form of heterodimer with the MSH5 protein (GeneCards database, http://www.genecards.org/cgi-bin/). MRE11 is a component of the MRN complex, which plays a central role in the repair of DNA double-strand breaks, recombination, and maintenance of telomeres integrity. MRE11 provides endonuclease activity on single-strand DNA and exonuclease activity on double-strand DNA (UniProt database, http://www.uniprot.org/). BRCA2 (see below information on BRCA1) participates in DNA recombination repair, due to assembling RAD51 on single-strand DNA, and also in the replication-dependent DNA double-strand breaks repair process (UniProt database). In meiosis, the process of silencing also takes place. Chromatin silencing, or suppression of gene expression, is an epigenetic process of gene regulation. Silencing can take place both at the transcription level and at the post-transcriptional level. Transcriptional silencing of unpaired chromatin regions (MSUC — meiotic silencing of unsynapsed chromatin) occurs by modifying histones, as a result of which the corresponding chromatin regions become inaccessible to RNA polymerase and transcription factors. This process occurs in meiosis in both females and males [15]. The MSUC process probably appeared initially in the course of the evolution as a selective mechanism for excluding cells with synaptic defects from the meiosis process. In meiosis, the need for silencing appears primarily in species with heteromorphic sex chromosomes, which remain unpaired for the most part [3]. In order for the cell to go through the control point and not react to unpaired DNA so that transcription from these DNA sections does not occur, the silencing mechanism (meiotic sex chromosome inactivation, MSCI) is turned on [13]. At the same time, autosomes remain transcriptionally active [4], including those regions where meiosis-specific genes are located. Morphological manifestation of the inactivation of chromosomes is the condensation of chromatin as a Barr body is forming (X-Y). Meiotic silencing has been found in different lines of the development of eukaryotes: in fungi, worms, insects, birds and mammals. The mechanism of silencing is common for these eukaryotic groups, but the details of the process may differ [4]. The molecular markers of the MSUC and MSCI processes are a large group of proteins involved in the repair of damaged DNA and chromatin remodeling factors [11], [18]. In particular, these are BRCA1, ATR, SUMO-1, XMR [8]. One of the main markers of transcriptional silencing of chromatin is phosphorylated histone H2AX (γH2AX), which is detected on asynaptiс chromosome sites, beginning with the leptotene stage up to the late diplotene, if the chromosome synapsis is not complete [16], [8]. The common ancestor of all eukaryotes had the H2AX histone gene. Nevertheless, the corresponding protein is absent in the yeast species S. pombe and S. cerevisiae, in many plants, as well as in insects (Drosophila, bee) and some mammals (GeneCards). The BRCA1 enzyme (E3 ubiquitin-protein ligase) has a broad spectrum of effect. It plays a central role in the process of DNA damage repair. In the case of MSCI, this enzyme attracts ATR kinase to the chromatin of sex chromosomes, which phosphorylates the H2AX histone and triggers the silencing process [14], (GeneCards). ATR is serine-threonine protein kinase (it is also ataxia-telangiectasia and Rad3-related protein). It phosphorylates the amino acid serine-139 in the specific meiotic histone H2AX/H2AFX at DNA damage sites, regulating the cell response to DNA damage. Orthologs of this protein are not found in some insects, fungi, plants and mammals (GeneCards). SUMO1 (small ubiquitin-related modifier 1, synonyms: GAPmodifying protein 1, ubiquitin-like protein SMT3C, etc.) works in the complex with other proteins. It takes part in such cellular processes as nuclear transport, DNA replication and repair, and others. Orthologs of this protein are absent in a number of plants and fungi, in insects (Drosophila, bee), and in some mammals (GeneCards). The main objective of this study was to analyze the conservation of key meiosis proteins using bioinformatics methods. Estimates published in the literature of a greater or lesser degree of evolutionary conservation of meiosis-specific proteins are qualitative and quantitative estimates based on different criteria for comparing proteins [12], [2], [17]. The study of recombination enzymes and silencing proteins in many laboratories continues until now, as evidenced at least by the materials of the latest international conference on meiosis (EMBO Conference on Meiosis 2017, 27th August — 1st September 2017, Hvar, Croatia). The conservation of enzymes has also been studied before (see, for example, the reviews of [1], [9]). However, for the first time we are examining a large group of proteins using a set of bioinformatics methods, which nobody has done before us.
2. System and methods
We have compared amino acid sequences of two protein groups. The first group consisted of 35 orthologs of enzymes involved in the formation of double DNA breaks (MRE11) and in the repair of chromatin damage in meiosis (MLH1, MSH4, BRCA1, BRCA2). The second group included 26 proteins taking part in chromatin silencing in meiosis (BRCA1 also takes part in this process). We have investigated SUMO1 orthologs (Small ubiquitin-related modifier 1 with a length of about 100 amino acid residues, a.a.), ATR (serine-threonine protein kinase with a size of about 2600 a.a.) and H2AX histone with a length of 130- 150 a.a. The indicated proteins were analyzed in eight model eukaryotes: fungi Saccharomyces cerevisiae and Schizosaccharomyces pombe, plant Arabidopsis thaliana, nematode Caenorhabditis elegans, insect Drosophila melanogaster, fish Danio rerio, mouse Mus musculus, and human being Homo sapiens. In case of BRCA1 and BRCA2 proteins, there were additionally analyzed the proteins of cocq Gallus gallus, lizard Anolis carolinensis, frog Xenopus laevis. Histone H2AX is absent in the proteomes of Drosophila and yeast S. pombe, so, we compared the corresponding proteins of other objects (histones of the frog Lithobates catesbeiana, insect Lygus hesperus, amoeba Dictyostelium fasciculatum, fungus Beauveria bassiana). In Arabidopsis, two histones were taken from different databases. Protein length was 130-150 a.a. (in the fungus, it was almost 500 a.a.). In the figures, the species are indicated either by the first letters of generic and species names, or abbreviated as in the UniProt database. Amino-acid sequences of the proteins were found in data bases UniProtKB/TrEMBL (http://www.uniprot.org/), NCBI (http://www.ncbi.nlm.nih.gov/guide/), and GeneCards (http://www.genecards.org/cgi-bin/). In case of the MSH4 enzyme of fish D. rerio and H2AX histone of Arabidopsis, these databases were contradictory to each other and we investigated two proteins of the same name (potential orthologs). For the BRCA2 protein of Arabidopsis, the UniProt database contains two homologues — A and B, as well as a protein similar to homologue A. We investigated the protein, which is called homologue A. In the absence of any protein, the corresponding proteins of other objects were studied in the model object. The presence of conserved functional domains was detected using the CDART program (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi?), and the set and sequence of conserved amino acid motifs — using the MEME program (http: // meme.nbcr.net/meme/tools/meme). To determine the secondary structure (the probability of forming an alpha-helical configuration), the COILS program (http://www.ch.embnet.org/software/COILS_form.html) was used, the isoelectric points of proteins (pI) were detected using Compute pI / Mw tool (http://web.expasy.org/compute_pi/). The severity of alpha helices (probability of formation from 0 to 1), the number and length of spiral fragments were evaluated. To evaluate the phylogenetic distance between orthologs, phylogenetic trees were constructed using the COBALT program (https://www.ncbi.nlm.nih.gov/tools/cobalt/cobalt.cgi?CMD=Web) in accordance with the Fast Minimum Evolution method. Evolutionary distance (be Grishin's method) between two sequences modeled as expected fraction of amino acid substitutions per site given the fraction of mismatched amino acids in the aligned region. For clarity, we measured the distance between the proteins of Arabidopsis and human because they are quite far from each other and present on all the trees that we have constructed.
2.1. Algorithm
First, we identified the domain structure of the supposed orthologs, which we selected from different databases (CDART program). If any protein did not have domains characteristic of other proteins of this family, it was discarded and was not further analyzed. At the next stage, conserved amino acid motifs (MEME) were detected. The secondary protein structure (COILS) and the isoelectric point (pI) were then analyzed. Based on all the data, a conclusion was made about the degree of conservation of each protein studied.
2.2. Implementation
The complex approach that we use has been tested for different groups of proteins, it is informative and can be used in the analysis of comparative conservation of orthologs in different lines of the development of eukaryotes.
3. Results
At the first stage, four parameters of conservation were analyzed for meiotic enzymes: MRE11, MLH1, MSH4, BRCA1, BRCA2. Orthologs of these proteins were analyzed in eight model species of eukaryotes, including humans. We note at once that one of the parameters — the formation of an alpha-helical secondary structure — turned out to be uninformative, because neither protein from both groups had uniformity in this characteristic. Thus, in case of BRCA1, the human protein has two helical regions, frog protein has three ones, and mouse protein has only one small helical fragment. In case of the MRE11 enzyme, vertebrate and yeast S. pombe (!) proteins have one expressed site of the alpha-helical configuration, Drosophila protein has one small helical fragment, and the proteins of nematode, Arabidopsis and yeast S. cerevisiae do not have any helical sites at all. Approximately the same picture is typical for other proteins we studied. The most conserved of the five studied meiotic enzymes was MLH1 (the length of the protein is from 650 to 750 a.a.). Its domain organization was the same for all studied species: a large domain MutL (DNA mismatch repair ATPase), which includes two smaller domains: HATPase (Histidine kinase-like ATPase) and MutL_Trans_MLH1 (transducer domain). By the set and arrangement of conserved amino acid motifs, the proteins are also similar (Figure 1), except that some motifs are absent in yeast (YEAST, SCHPO) and nematode (CAEEL). There are four regions of the greatest conservation: N-terminus fragment, two sites in the middle of the molecule and a small site at the C-terminus. The isoelectric points (pI) of the studied orthologs MLH1 were in a narrow interval (5.40-6.08), except for the Drosophila protein (8.08). The phylogenetic distance between the proteins of Arabidopsis and human measured on the corresponding phylogenetic tree constitutes approximately one conventional unit.
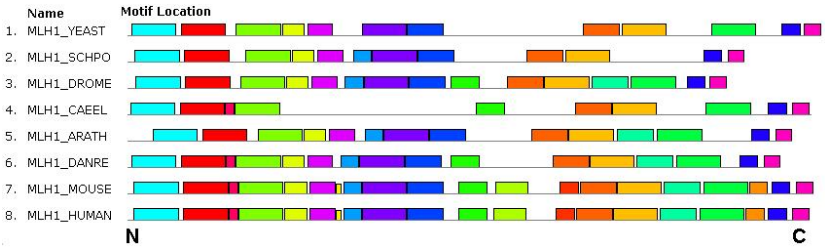
Figure 1 – The location of conserved amino acid motifs in MLH1 protein molecules of the human (HUMAN), mouse (MOUSE), fish (DANRE), nematode (CAEEL), Drosophila (DROME), plant (ARATH) and yeast (YEAST and SCHPO). The common motifs are shown by rectangles of the same colors and the same sizes. The N- and C-terminus of the protein molecule are indicated.
A little less conserved was the MRE11 enzyme (protein length about 700 a.a.). Its orthologs had a similar set of motifs at the N-terminus and in the middle part of the molecule (in nematode and Drosophila, this fragment of the protein did not have all the motifs). The C-terminus fragments of proteins differed in all but the mouse and human. The protein contains one functional domain from the family Mre11_DNA_bind, occupying the first half of the molecule. The pI range was also rather narrow (5.35- 5.70), with the exception of the Arabidopsis protein (6.21). The phylogenetic distance between MRE11 of Arabidopsis and human measured on the phylogenetic tree constitutes approximately 1.3 units. Even less conserved was the MSH4 protein (protein length was about 1000 a.a.). In Drosophila and yeast S. pombe, this enzyme is not present; in D. rerio fish, two proteins from different databases were examined. Although almost all studied orthologs have one large MutS domain from the MutS_III superfamily, a set of conserved motifs was similar only in proteins of vertebrate animals. Small motifs in the N-terminus and middle parts of the molecule and a small motif closer to the C-terminus were common to all studied orthologs. The isoelectric points (pI) were also more scattered (from 6.28 to 8.47). The least conserved were BRCA1 and BRCA2 proteins. They are present only in vertebrates, and not in all (in some databases, there were also annotated orthologs for Arabidopsis). In the BRCA1 protein, there were revealed small functional domains at the ends of the molecule. The length of orthologs varied greatly — from 900 to 1800 a.a. Similar motifs were also observed only at the N- and C-terminus (Figure 2). The protein of Arabidopsis differed from proteins of vertebrates under both characteristics. The isoelectric points of BRCA1 in vertebrates turned out to be quite similar (5.29-5.73), while in Arabidopsis it was 9.18. This may indicate that Arabidopsis does not really have BRCA1. The phylogenetic distance between BRCA1 proteins of Arabidopsis and human measured on the corresponding phylogenetic tree constitutes approximately 2.5 units.
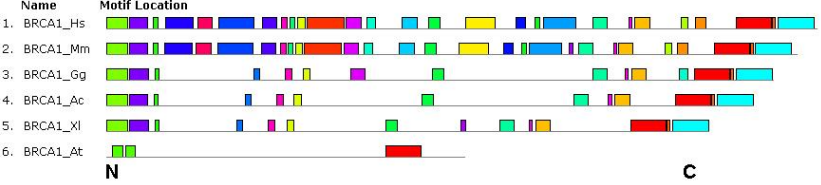
Figure 2 – The location of conserved amino acid motifs in human (Hs), mouse (Mm), cocq (Gg), lizard (Ac), frog (Xl) and plant (At) BRCA1 protein molecules.
The common motifs are shown by rectangles of the same colors and the same sizes. The N- and C-terminus of the protein molecule are indicated.
The BRCA2 protein is absent in yeast and nematode, it is annotated in Drosophila but smaller in size (does not carry domains, therefore it was excluded from analysis). In Arabidopsis, there are annotated three similar proteins, we analyzed one. Cocq and lizard proteins were also taken in the analysis. The length of orthologs varied approximately from 1150 to 3400 a.a. The orthologs that we studied carry functional domains only in the C-terminus fragment (in the lizard, the same domains are in the central part of the molecule). Not all domains have been revealed in Arabidopsis. We examined the C-terminus fragment of the protein in terms of the presence of common motifs. Even within the subtype of vertebrates, the set and location of conserved motifs slightly vary. The range of isoelectric points is quite wide (6.02-8.77). At the second stage, we analyzed the conservation of the key proteins of chromatin silencing in meiosis: SUMO1, ATR and H2AX histone. The isoelectric points of the orthologs of two of the three proteins were in rather narrow intervals: the SUMO1 protein turned out to be very acidic (pI from 4.62 to 5.34), and the H2AX histone, in contrast, was alkaline (pI interval 9.83 to 10.83). Standing out from others is the only histone of the fungus Beauveria bassiana (pI=5,15).. We assumed that this protein is not a true H2AX histone. For ATR, the range of pI values was slightly wider, from 6.62 to 8.54. Thus, according to this parameter, the ATR protein is less conserved than the other two ones. This conclusion was confirmed in the study of functional domains and conserved amino acid motifs of these proteins.
In most species studied, H2AX histone has one large domain from the H2 superfamily, occupying almost the entire length of the molecule (length of the protein is 130-150 a.a.). It is slightly modified in the amoeba and insect. The fungus Beauveria bassiana has only a fragment of such a domain at the C-terminus of the protein. Apparently, the histone, annotated for this species as H2AX, is not that. The set and sequence of conserved motifs were almost identical for all species except the fungus (Figure 3). The previous conclusion was confirmed. The H2AX histone was the most conserved of the three proteins studied. It is noted in the GeneCards database that a corresponding gene was present in the ancestors of all eukaryotes. As for conservation, this histone can be compared to the MLH1 enzyme.
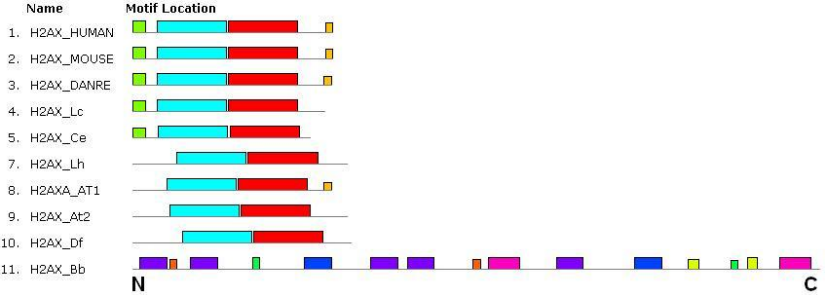
Figure 3 – The set and sequence of conserved motifs in H2AX histones of the human (HUMAN), mouse (MOUSE), fish (DANRE), frog (Lc), nematode (Ce), insect (Lh), Arabidopsis (At1, At2), amoeba (Df) and fungus (Bb).
The N- and C-terminus of the protein molecules are shown. The common motifs are indicated by rectangles of the same colors and sizes.
The SUMO1 protein, according to the GeneCards website, is absent in Arabidopsis, Drosophila and S. pombe yeast. However, in other databases, the proteins for Arabidopsis and yeast are annotated, and we included them in the analysis. In molecules of this small protein (about 100 a.a.) one domain is found that occupies almost the entire molecule. Moreover, in the yeast S. pombe, another domain was present, although belonging to the same family. In terms of the set of conserved motifs, all the studied proteins were more similar. Only N-terminus sections of molecules were different. Thus, SUMO1 is slightly less conserved than histone and the MLH1 enzyme. Let us now proceed to the analysis of the ATR protein also known as ataxia-telangiectasia and Rad3-related protein. Its orthologs about 2600 a.a. long differed already at the level of functional domains (different number of domains and their different sizes). Only representatives of vertebrates, as well as Drosophila have been found similar in this respect (all of them have 4 domains closer to the C-terminus of the molecule). In Arabidopsis, there were revealed three of the four domains; in the nematode and yeast — one in each, but they are different. In terms of the set and sequence of conserved amino acid motifs, the N-terminus fragments of ATR proteins are similar only in vertebrates, the middle part of the molecule has no similarity, and only C-terminus motifs are present in all studied species, although their sets slightly differ (Figure 4). Thus, the ATR protein is the least conserved of the three studied silencing proteins of the meiotic chromatin.
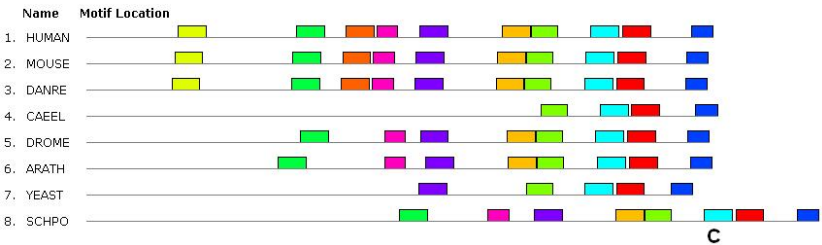
Figure 4 – The set and sequence of conserved motifs in ATR proteins of the human (HUMAN), mouse (MOUSE), fish (DANRE), nematode (CAEEL), Drosophila (DROME), plant (ARATH), yeast S. cerevisiae (YEAST) and S. pombe (SCHPO).
A C-terminus fragment of protein molecules with a length of about 1000 a.a. is shown. The common motifs are indicated by rectangles of the same colors and sizes.
In conclusion, the practical significance of the results obtained should be noted. In laboratories where immunocytochemical studies are carried out on mammalian, reptilian, plant and fungal cells, it is important to select antibodies suitable for different objects (species), since these are costly reagents. Knowing the location and size of conserved motifs helps make this choice, because antibodies are selected based on the similarity of amino acid sequences in ortholog proteins.
4. Discussion
The conservation of five meiosis enzymes was studied: MRE11, MLH1, MSH4, BRCA1, BRCA2. The former is involved in the formation of double DNA breaks, including mitosis, MLH1 and MSH4 (meiosis-specific) and BRCA1-2 are involved in the repair of DNA damage. The probability of the formation of an alpha-helical structure is the parameter that we traditionally investigate when assessing the conservation of these or other meiotic proteins. However, the importance of this structure is not always high. It is critical for structural proteins, for example, for those forming transverse filaments of the synaptonemal complex. In representatives of different lines of development of eukaryotes, such proteins do not have homology of the primary sequence, but they have an extended alpha-helix in the central part of the molecule, which allows them to form rod-like structures [2]. Shugoshins — the centrosome cohesion protectors in meiosis — have a diagnostic alpha-helical region in the N-end of the molecule, and in other parts of the molecule the number and severity of the helices may be different [5]. Apparently, this parameter is not critical for recombination enzymes and silencing factors. The relatively high conservation of the protein for correcting mispaired bases (mismatch repair) MLH1 is not surprising, since this protein performs the same function in all organisms: control over compliance with the rules of complementarity of nucleotides in the structure of the DNA double helix [1]. It also participates in the "ripening" of reciprocal exchange events [9]. We have previously shown that the SPO11 exonuclease, in the ensemble with which the MRE11 enzyme works, is not conserved [6]. However, its partner, the MRE11 protein, involved in the formation of the 3'-single-strand end of DNA, proved to be quite conserved, especially in the area of the functional domain (the first two-thirds of the molecule are conserved). The MSH4 enzyme is a partner of MLH1, but not in the repair of mismatched bases, but in the repair of early DNA breaks, while contacting with the proteins of the synaptonemal complex [9]. Apparently, the requirements for its conservation are lower. Only the enzymes of vertebrate animals are rather similar, in spite of the fact that identical functional domains are revealed in all orthologs. As for the somatic repair enzymes BRCA1 and BRCA2, working in meiosis (detection of demage, transmission of a signal about it, attraction of repair enzymes, [9], they appear to be present only in vertebrates and show little similarity even within this subtype. They are similar only in the C-terminus part of the molecule. The H2AX histone plays a central role in regulating transcription, in DNA repair and replication [16], [8], GeneCards. The phosphorylated variant of this histone (γH2AX) is the first marker of DNA break and is necessary to attract other proteins involved in DNA repair. It turned out to be the most conserved of the silencing proteins we studied, apparently because it directly contacts DNA and a lot of partner proteins. As for conservation, it can be compared with the MLH1 enzyme. A little less conserved than histone and the MLH1 enzyme was the small protein SUMO1 (small ubiquitin-related modifier 1), which takes part in such cellular processes as nuclear transport, DNA replication and repair, mitosis and signal transduction (GeneCards). In contrast to these proteins, the ATR kinase (serine-threonine protein kinase, the same as ataxia-telangiectasia), turned out to be the least conserved protein among all studied by us: similarity was observed only in the C-terminus fragments of orthologs proteins. For the sake of justice, it should be noted that the size of this protein is very large (about 2600 a.a.), so, it was difficult to expect the similarity of the entire amino acid sequence in the orthologs studied by us. Orthologs of this protein are not found in some insects, fungi, plants and mammals [GeneCards]. Thus, the meiosis proteins studied by us in terms of the degree of conservation can be arranged as follows: MLH1, H2AX > SUMO1 > MRE11 > MSH4 > BRCA1, BRCA2, ATR. The phylogenetic distances between the proteins of Arabidopsis and human for MLH1, MRE11 and BRCA1 proteins located at the beginning, middle and end of this line are in line with our conclusions (one conventional unit, 1.3 units and 2.5 units, respectively).
Supplementary materials
Does not apply.
Funding
This work was supported by Russian Foundation for Basic Research [project № 16-04-01447 а] and the State Assignment [Contract № 0112-2016-0008] with the use of the equipment of the SC "Genetic polymorphism" of Department of Biological Sciences RAS.
Acknowledgement
Does not apply.
Conflict of Interest
None declared.
References
Anuradha, S. Saccharomyces cerevisiae Hop1 zinc finger motif is the minimal region required for its function in vitro [text] / Anuradha, S., Muniyappa, K. // J. Biol. Chem. – 2004. – V. 279. – P. 28961–28969.
Bogdanov, Yu.F. Similarity of the domain structure of proteins as a basis for the conservation of meiosis [text] / Bogdanov, Yu.F., Grishaeva, T.M., Dadashev, S.Ya. // Intern. Rev. Cytol. – 2007. – V. 257. – P. 84–142.
Burgoyne, P.S. Genetic homology and crossing over in the X and Y chromosomes of Mammals [text] / Burgoyne, P.S. // Hum. Genet. – 1982. – V. 61. – № 2. – P. 85–90.
Daish, T.J. Lack of sex chromosome specific meiotic silencing in platypus reveals origin of MSCI in therian mammals [text] / Daish, T.J. // BMC Biology. – 2015. – V. 13. – P. 106–118.
Grishaeva T.M. Bioinformatical analysis of Eukaryotic shugoshins revealed meiosis-specific features of vertebrate shugoshins [text] / Grishaeva T.M., Kulichenko D.А., Bogdanov Y.F. // PeerJ. – 2016. – V. 4. – P. e2736.
Grishaeva, T.M. Evolutionary conservation of recombination proteins and variability of meiosis-specific proteins of chromosomes [text] / Grishaeva, T.M., Bogdanov, Y.F. // Russian J. Genetics. – 2017. – V. 53. – № 5. – P. 542–550.
Hunter, N. The single-end invasion: An asymmetric intermediate at the double-strand break to double-holliday junction transition of meiotic recombination [text] / Hunter, N., Kleckner, N. // Cell. – 2001. – V. 106. – P. 59¬–70.
Manterola, M. A high incidence of meiotic silencing of unsynapsed chromatin is not associated with substantial pachytene loss in heterozygous male mice carrying multiple simple robertsonian translocations [text] / Manterola, M., Page, J., Vasco, C., Berríos, S., Parra, M., Viera, A., Rufas, J., Zuccotti, M., Garagna, S., Fernández–Donoso, R. // PLoS Genet. – 2009. – V. 8. – P. 1–14.
Marcon, E. The evolution of meiosis: recruitment and modification of somatic DNA-repair proteins [text] / Marcon, E., Moens, P.B. // Bioessays. – 2005. – V. 27. – P. 795–808.
Priming, M. The core meiotic transcriptome of budding yeast [text] / Priming, M., Williams, R., Winzeler, E.A., Tevzadze, G.G., Conway, A.R., Hwang, S.Y., Davis, R.W., Esposito, R.E. // Nature Genet. – 2000. – V. 26. – P. 415–423.
Schimenti, J. Synapsis or silence [text] / Schimenti, J. // Nat. Genet. – 2005. – V. 37. – № 1. – P. 11–13.
Stassen, N.Y. Isolation and characterization of rad51 orthologs from Coprinus cinereus and Lycopersicon esculentum, and phylogenetic analysis of eukaryotic recA homologs [text] / Stassen, N.Y., Logsdon, J.M. Jr, Vora, G.J. Offenberg, H.H., Palmer, J.D., Zolan, M.E. // Curr. Genet. – 1997. – V. 31. – P. 144–157.
Turner, J.M. Meiotic sex chromosome inactivation [text] / Turner, J.M. // Development. – 2007. – V. 34. – № 10. – P. 1823–1831.
Turner, J.M. BRCA1, histone H2AX phosphorylation, and male meiotic sex chromosome inactivation [text] / Turner, J.M., Aprelikova, O., Xu, X., Wang, R., Kim, S., Chandramouli, G.V.R., Barrett, J.C., Burgoyne, P.S., Deng, C.-X. // Current Biology. – 2004. – V. 14. – P. 2135–2142.
Turner, J.M. Silencing of unsynapsed meiotic chromosomes in the mouse [text] / Turner, J.M., Mahadevaiah, S.K., Fernandez-Capetillo, O., Nussenzweig, A., Xu, X., Deng, C.X., Burgoyne, P.S. // Nat. Genet. – 2005. – V. 37 – № 1. P. 41–47.
Turner, J.M.A. Pachytene asynapsis drives meiotic sex chromosome inactivation and leads to substantial postmeiotic repression in spermatids [text] / Turner, J.M.A., Mahadevaiah, S.K., Ellis, P.J.I., Mitchell, M.J., Burgoyne, P.S. // Develop. Cell. – 2006. – V. 10. – № 4. – P. 521–529.
Waldman, B-A.H. Expression and chromosomal organization of mouse meiotic genes [text] / Waldman, B-A.H., Shahar, I., Yitzchak, A., Mehr, R., Don, J. // Mol. Reprod. Develop. – 2010. – V. 77. – P. 241–248.
Zamudio, N.M. Epigenetic regulation in male germ cells [text] / Zamudio, N.M., Chong, S., O'Bryan, M.K. // Reproduction. – 2008. – V. 136. – № 2. – P. 131–146.
References
Anuradha, S. Saccharomyces cerevisiae Hop1 zinc finger motif is the minimal region required for its function in vitro [text] / Anuradha, S., Muniyappa, K. // J. Biol. Chem. – 2004. – V. 279. – P. 28961–28969.
Bogdanov, Yu.F. Similarity of the domain structure of proteins as a basis for the conservation of meiosis [text] / Bogdanov, Yu.F., Grishaeva, T.M., Dadashev, S.Ya. // Intern. Rev. Cytol. – 2007. – V. 257. – P. 84–142.
Burgoyne, P.S. Genetic homology and crossing over in the X and Y chromosomes of Mammals [text] / Burgoyne, P.S. // Hum. Genet. – 1982. – V. 61. – № 2. – P. 85–90.
Daish, T.J. Lack of sex chromosome specific meiotic silencing in platypus reveals origin of MSCI in therian mammals [text] / Daish, T.J. // BMC Biology. – 2015. – V. 13. – P. 106–118.
Grishaeva T.M. Bioinformatical analysis of Eukaryotic shugoshins revealed meiosis-specific features of vertebrate shugoshins [text] / Grishaeva T.M., Kulichenko D.А., Bogdanov Y.F. // PeerJ. – 2016. – V. 4. – P. e2736.
Grishaeva, T.M. Evolutionary conservation of recombination proteins and variability of meiosis-specific proteins of chromosomes [text] / Grishaeva, T.M., Bogdanov, Y.F. // Russian J. Genetics. – 2017. – V. 53. – № 5. – P. 542–550.
Hunter, N. The single-end invasion: An asymmetric intermediate at the double-strand break to double-holliday junction transition of meiotic recombination [text] / Hunter, N., Kleckner, N. // Cell. – 2001. – V. 106. – P. 59¬–70.
Manterola, M. A high incidence of meiotic silencing of unsynapsed chromatin is not associated with substantial pachytene loss in heterozygous male mice carrying multiple simple robertsonian translocations [text] / Manterola, M., Page, J., Vasco, C., Berríos, S., Parra, M., Viera, A., Rufas, J., Zuccotti, M., Garagna, S., Fernández–Donoso, R. // PLoS Genet. – 2009. – V. 8. – P. 1–14.
Marcon, E. The evolution of meiosis: recruitment and modification of somatic DNA-repair proteins [text] / Marcon, E., Moens, P.B. // Bioessays. – 2005. – V. 27. – P. 795–808.
Priming, M. The core meiotic transcriptome of budding yeast [text] / Priming, M., Williams, R., Winzeler, E.A., Tevzadze, G.G., Conway, A.R., Hwang, S.Y., Davis, R.W., Esposito, R.E. // Nature Genet. – 2000. – V. 26. – P. 415–423.
Schimenti, J. Synapsis or silence [text] / Schimenti, J. // Nat. Genet. – 2005. – V. 37. – № 1. – P. 11–13.
Stassen, N.Y. Isolation and characterization of rad51 orthologs from Coprinus cinereus and Lycopersicon esculentum, and phylogenetic analysis of eukaryotic recA homologs [text] / Stassen, N.Y., Logsdon, J.M. Jr, Vora, G.J. Offenberg, H.H., Palmer, J.D., Zolan, M.E. // Curr. Genet. – 1997. – V. 31. – P. 144–157.
Turner, J.M. Meiotic sex chromosome inactivation [text] / Turner, J.M. // Development. – 2007. – V. 34. – № 10. – P. 1823–1831.
Turner, J.M. BRCA1, histone H2AX phosphorylation, and male meiotic sex chromosome inactivation [text] / Turner, J.M., Aprelikova, O., Xu, X., Wang, R., Kim, S., Chandramouli, G.V.R., Barrett, J.C., Burgoyne, P.S., Deng, C.-X. // Current Biology. – 2004. – V. 14. – P. 2135–2142.
Turner, J.M. Silencing of unsynapsed meiotic chromosomes in the mouse [text] / Turner, J.M., Mahadevaiah, S.K., Fernandez-Capetillo, O., Nussenzweig, A., Xu, X., Deng, C.X., Burgoyne, P.S. // Nat. Genet. – 2005. – V. 37 – № 1. P. 41–47.
Turner, J.M.A. Pachytene asynapsis drives meiotic sex chromosome inactivation and leads to substantial postmeiotic repression in spermatids [text] / Turner, J.M.A., Mahadevaiah, S.K., Ellis, P.J.I., Mitchell, M.J., Burgoyne, P.S. // Develop. Cell. – 2006. – V. 10. – № 4. – P. 521–529.
Waldman, B-A.H. Expression and chromosomal organization of mouse meiotic genes [text] / Waldman, B-A.H., Shahar, I., Yitzchak, A., Mehr, R., Don, J. // Mol. Reprod. Develop. – 2010. – V. 77. – P. 241–248.
Zamudio, N.M. Epigenetic regulation in male germ cells [text] / Zamudio, N.M., Chong, S., O'Bryan, M.K. // Reproduction. – 2008. – V. 136. – № 2. – P. 131–146.
