Return to article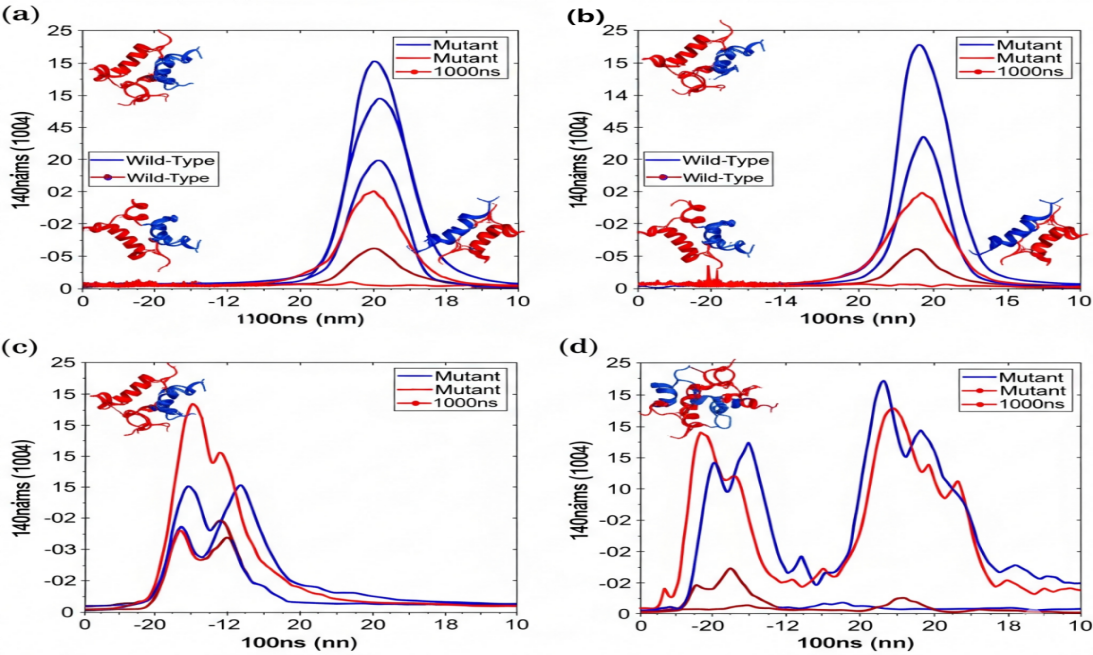
Преодоление разрыва между генотипом и фенотипом: интеграция полногеномных ассоциативных исследований и вычислительной структурной биологии для расшифровки механизмов неправильного фолдинга белков, вызванного однонуклеотидными полиморфизмами, при заболеваниях человека
Molecular Dynamics Simulation Results for a Destabilizing Mutation

Wild-Type (blue) and Mutant (red) are compared over 100ns MD simulation via a set of graphs: a) Root Mean Square Deviation (RMSD). Mutant is characterized by higher and more varied RMSD, thus indicating the structural instability; b) Root Mean Square Fluctuation (RMSF). The increased fluctuations in the Mutant can be found at certain functional regions (e.g., binding loops 1-3); с) Solvent Accessible Surface Area (SASA). The Mutant shows a higher SASA that suggests the partial unfolding and the hydrophobic core being exposed; d) Number of Hydrogen Bonds. The stable intra molecular H-bonds in the Mutant are fewer
