Return to article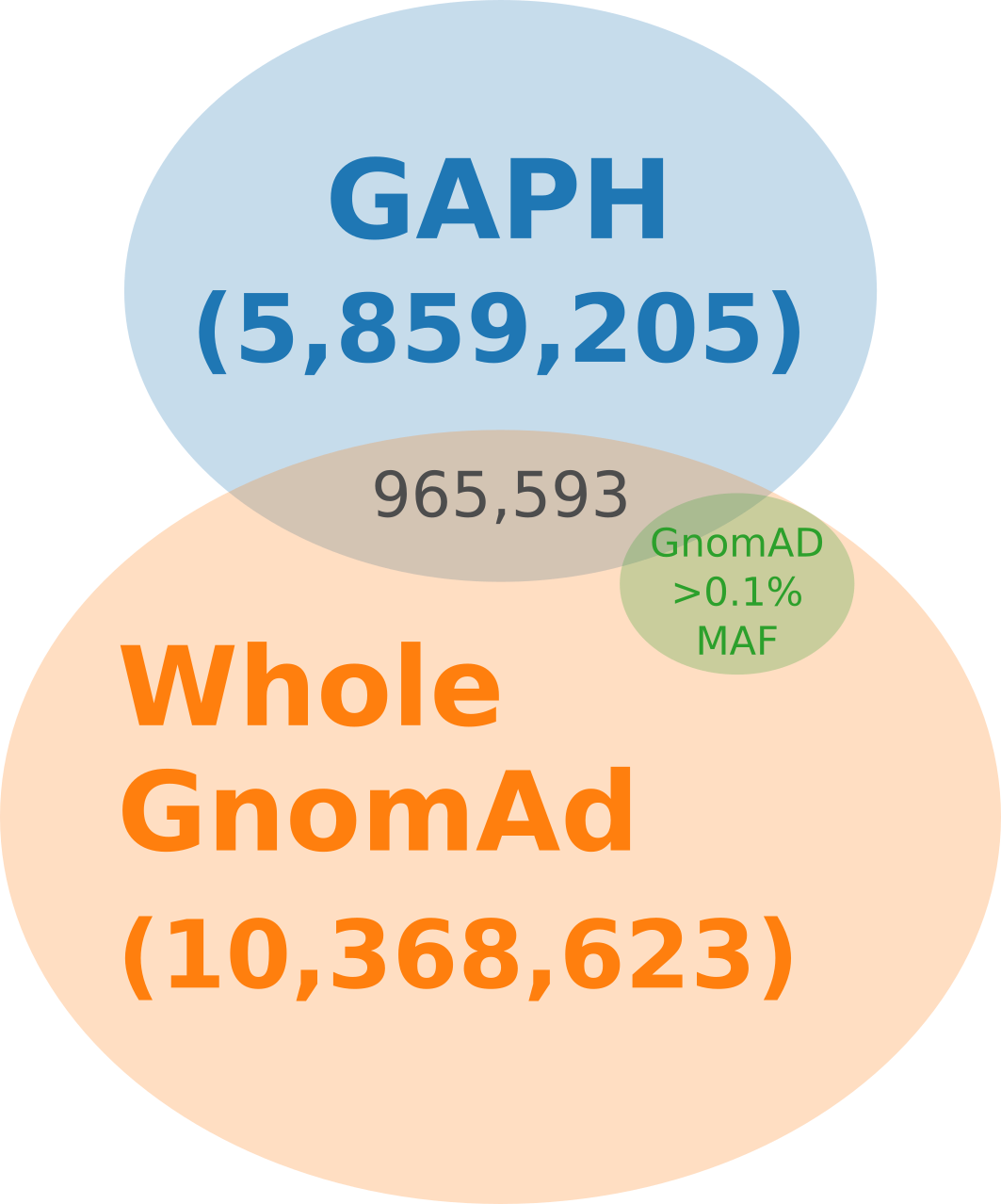
ВЫРАВНИВАНИЕ ПСЕВДОРИДОВ ГОМОЛОГИЧНЫХ ПОСЛЕДОВАТЕЛЬНОСТЕЙ В ОПРЕДЕЛЕНИИ ПОТЕНЦИАЛЬНО ПЕРЕНОСИМЫХ ГЕНОМНЫХ ВАРИАНТОВ

Venn diagram illustrating the overlap between GAPH-resulted set, variants found in GnomAD v3.1.2, and subset of GnomAD variants with minor allele frequency (MAF) > 0.1%.
